Ultra-Deep Microbiome Prep: Host DNA Depletion for Accurate Analysis |
| 25 October 2019 |
|
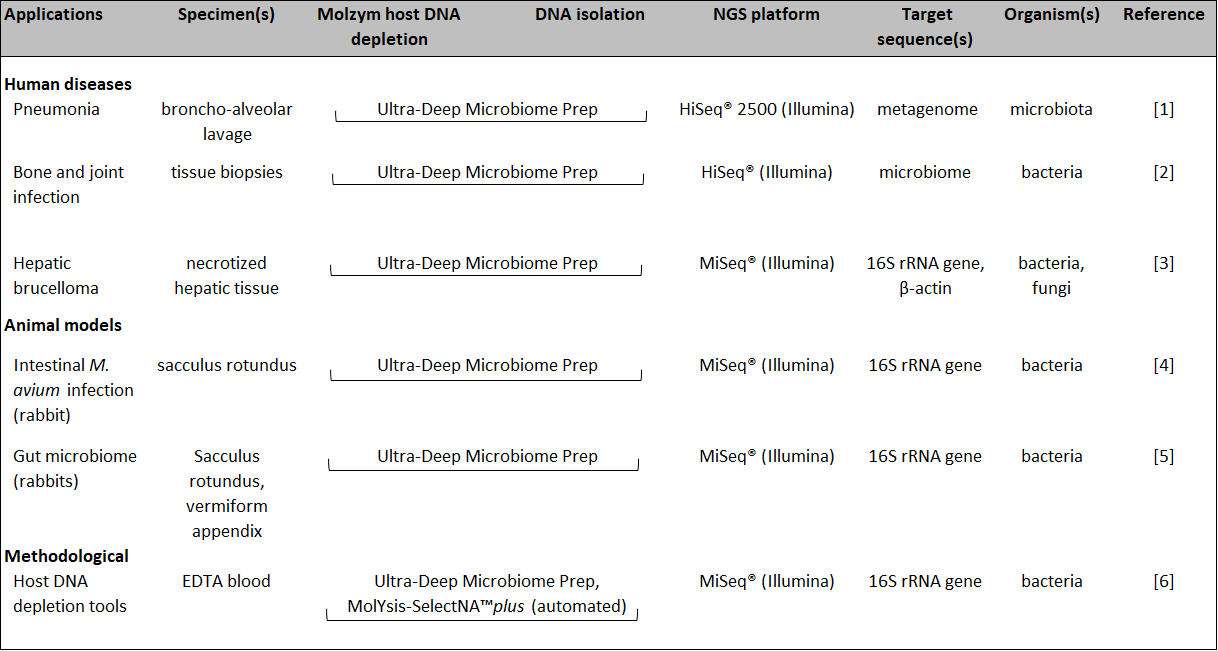
Host DNA is present at vast excess to the target microbial DNA in many sampling materials. This poses problems on amplification-based analysis using primers directed to microbial sequences. The reason is that at low bacterial or fungal loads primers tend to bind unspecifically to host DNA which lowers the sensitivity of detection and, in NGS analysis, generates poor reads of microbial origin and hence low depth of microbiome structures. Reagent-borne contamination by microbial DNA is another problem that can disturb the analysis of microorganisms endogenous in a sample. With its MolYsis™ technology, Molzym provides a solution to the depletion of host DNA under exclusion of reagent-borne contaminations. The technology is based on the enzymatic degradation of human or animal DNA followed by the isolation of thus enriched microbial DNA. All reagents and plastics of the Ultra-Deep Microbiome Prep kit are guaranteed free of microbial DNA over 40 PCR cycles. The kit allows the processing of liquid and tissue specimens. The isolated microbial DNA can be used with various systems of next generation sequencing microbiome analysis. Examples of applications using the Illumina® systems are summarized below, including the diagnosis of human and animal microbiomes in liquid and tissue specimens. |
 |
|
More information on the MolYsis™ technology for NGS analyses is given in a recent application note. |
| References |
| [1] Leo S et al. (2017) Int J Mol Sci 18(9):2011, doi:10.3390/ijms18092011. Go to blog post |
| [2] Ruppé et al. (2017) Scientific Reports volume 7, Article number: 7718, doi:10.1038/s41598-017-07546-5. |
| [3] Lazarevic V et al. (2018) Front Microbiol. 2018;9:1566. doi:10.3389/fmicb.2018.01566. |
| [4] Arrazuria R et al. (2016) Front Microbiol 7, 446. doi: 10.3389/fmicb.2016.00446. |
| [5] Arrazuria et al. (2018) Scientific Reports 8, 14103. doi:10.1038/s41598-018-32484-1. |
| [6] Edelmann A et al. (2018) Poster ECCMID 2018 #P0082. Go to blog post |

